A large part of the success of GPT-3.5 and GPT-4 is attributed to the fact that these models did undergo, in addition to pre-training and supervised instruction fine-tuning, a third phase of learning called reinforcement learning with human feedback. There are many posts on this, but unfortunately most of them fall short of explaining how the training method really works and stay a bit at the surface. If you want to know more – read on, we will close this gap today which also concludes this series on large language models. But buckle on, this is going to be a long post.
What is reinforcement learning?
Pretraining a language model using teacher forcing and fine-tuning using supervised training have one thing in common – a ground truth, i.e. a label, is known at training time. Suppose for instance we train a model using teacher forcing. We then feed the beginning of a sentence that is part of our training data into the model and, at position n, teach the model to predict the token at position n + 1. Thus we have a model output and a label, calculate a loss function that measures the difference between those two and use gradient descent to minimize the loss. Similarly, if we fine-tune on a set of question-instruction pairs, we again have a model output and a label that we can compare in exactly the same way.
However, language is more complicated than this. Humans do not only rate a reply to a question on the level of individual token, but also by characteristics that depend on the sentence as a whole – whether a reply is considered as on topic or not is not only a function of the individual token that appear in it.
So if we want to use human feedback to train our model, we will have to work with feedback, maybe in the form of some score, that is attached not to an individual token, but to the entire sequence of token that make up a reply. To be able to apply methods like gradient descent, we need to somehow propagate that score down the level of individual token, i.e. sampling steps. Put differently, we will need to deal with delayed feedback, i.e. constituents of the loss function that are not known after sampling an individual token but only at a later point in time when the entire sentence is complete. Fortunately, the framework of reinforcement learning gives us a few tools to do this.
To understand the formalism of reinforcement learning, let us take a look at an example. Suppose that you wanted to develop a robot that acts as an autonomous vacuum cleaner which is able to navigate through your apartment, clean as much of it as possible and eventually return to a docking station to re-charge its batteries. While moving through the apartment, the robot needs to constantly take decisions like moving forward or backward or rotating to change direction. However, sometimes the benefit of a decision is not fully apparent at the point in time when it has to be taken. If entering a new room, for instance, the robot might enter the room and try to clean it as well, but doing this might move it too far away from the charging station and it might run out of energy on the way back.
This is an example where the robot has to take a decision which is a trade off – do we want to achieve a short-term benefit by cleaning the additional room or do we consider the benefit of being recharged without manual intervention as more important and head back to the charging station? Even worse, the robot might not even know the surface area of the additional room and would have to learn by trial-and-error whether it can cover the room without running out of power or not.
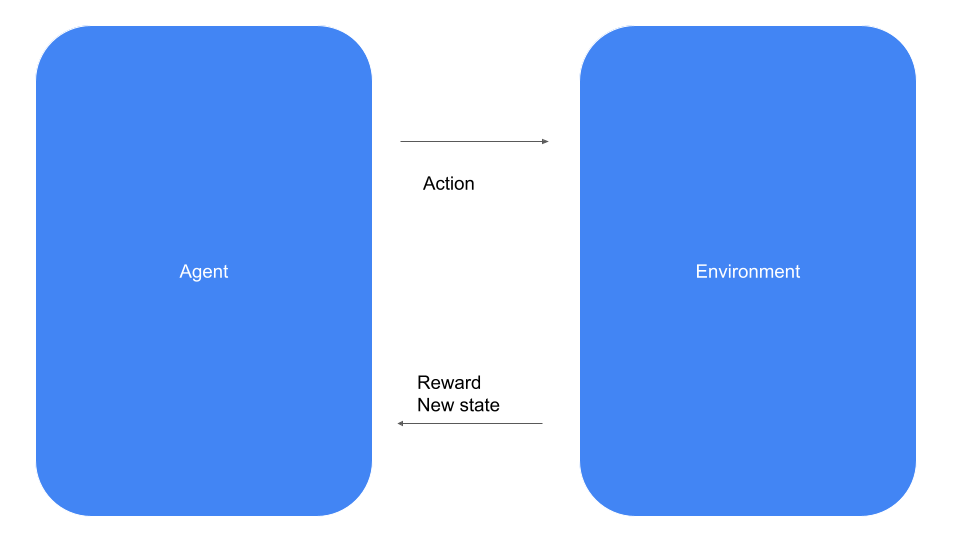
Let us formalize this situation a bit. Reinforcement learning is about an agent like our vacuum cleaning robot that operates in an environment – our apartment – and, at each point in time, has to select one out of a set of possible actions, like moving forward, backward, left or right. When selecting an action, the robot receives a feedback from the environment called a reward, which is simply a scalar value that signals the agent whether the decision it has taken is considered beneficial or not. In our example, for instance, we could give the robot a small positive reward for every square meter it has cleaned, a larger positive reward for reaching the charging station and a small negative reward, i.e. a penalty, when it bumps into a wall.
In every time step, the agent can select one the available actions. The environment will then provide the reward and, in addition, update the state of the system (which, in our case, could be the location of the agent in the apartment and the charging state of the battery). The agent then receives both, the reward and the new state, as an information and proceeds with the next time step.
Let us now look at two ways to formalize this setting – mathematics and code. In mathematical terms, a problem in reinforcement learning is given by the following items. First, there is a set of states, which we typically assume to be finite, and a set
of possible actions, which we assume to be finite as well. In addition, there is a set
of rewards.
The reward that the agent receives for a specific action and the next state depend on the current state and the action. We could now proceed and define this is a mapping
However, in many cases, this assignment has a probabilistic character, i.e. the reward and the next state are not fully deterministic. Therefore, the framework of a Markov decision process (MDP) that we use here describes rewards and next state via a probability distribution conditioned on the current state and the action, i.e. as
where St+1 is the state at time t + 1, St is the state at time t, At is the action taken at time t and Rt+1 is the reward that the agent receives at time t (this convention for the index of the reward might appear a bit unusual, but is in fact quite common in the field). Collectively, these conditional probabilities are called transition probabilities.
Note that we are making two crucial assumptions here. The first one is that the reward and next state depend only on the action and the current state, not on any previous state and not on the history of events so far (Markov property). This is rather a restriction when modelling a real world problem, not a restriction on the set of problems to which we can apply the framework, as we could simply redefine our state space to be the full history up to the current timestep. Second, we assume that the agent has access to the full state on which rewards and next states depend. There are also more general frameworks in which the agent only receives an observation that reflects a partial view on the actual state (this is the reason why some software packages use the term observation instead of state).
Let us now look at this from a different angle and see how this is modeled as an API. For that purpose, we will look at the API of the popular Gymnasium framework. Here, the central object is an environment that comes with two main methods – step and reset. The method reset resets the environment to an initial state and returns that state. The method step is more interesting. It accepts an action and returns a reward and the new state (called an observation in the Gym framework), exactly as indicated in the diagram above.
Thus an agent that uses the framework would essentially sit in a loop. In every step, it would pick an action that it wants to perform. It would then call the step method of the environment which would trigger the transition into a new state, depending on the action passed as argument. From the returned data, the agent would learn about the reward it has received as well as about the next state.
When would the loop end? This does of course depend on the problem at hand, but many problems are episodic, meaning that at some point in time, a terminal state is reached which is never left again and in which no further rewards are granted. The series of time steps until such a terminal state is reached is called an episode. We will see later than in applications to language modelling, an episode could be a turn in a dialogue or a sentence until an end-of-sentence token is sampled. However, there are also continuing tasks which continue potentially forever. An example could be a robot that learns to walk – there is no obvious terminal state in this task, and we would even encourage the robot to continue walking as long as possible.
Policies and value functions
We have now described a mathematical framework for states and actions. What we have not yet discussed, however, is how the agent actually takes decisions. Let us make the assumption that the agents decisions only depend on the state (and not, for instance, on the time step). The rule which assigns states to actions is called the policy of the agent. Again, we could model this in a deterministic way, i.e. as a function
but it turns out to be useful to also allow for stochastic policies. A policy is then again a probability distribution which describes the probability that the agent will take a certain action, conditioned on the current state. Thus, a policy is a conditional probability
Note that given a policy, we can calculate the probability to move into a certain state s’ starting from a state s as
giving us a classical Markov chain.
What makes a policy a good policy? Usually, the objective of the agent is to maximize the total reward. More precisely, for every time step t, the reward Rt+1 at this time step is a random variable of which we can take the expectation value. To measure how beneficial being in a certain state s is, we could now try to use the sum
of all these conditional expectation values as a measure for the total reward. Note that the conditioning simply means that in order to calculate the expectation values, we only consider trajectories that start at given state s.
However, it is not obvious that this sum converges (and it is of course easy to come up with examples where it does not). Therefore, one usually builds a discounting factor into this equation and defines the value of a state s to be the conditional expectation value
Note that the discounting factor plays two roles in this definition. First, it makes sure that under mild conditions, for instance a bound on the rewards, the sum is finite. Second, the value of a reward received early in the trajectory is higher than the value of the same reward at a later time step, so the discounting encourages the agent to collect rewards as early as possible. Whether this is reasonable for the problem at hand is a matter of modelling, and in real implementations, we often see discounting factors close to one.
The mapping that assigns to each state s the value v(s) defined above is called the value function. Note that the expectations are taken with respect to the state transition probabilities given by the policy , and in fact the value function depends on the policy. In general, the objective of the agent will be to find a policy which maximizes the value function for all states or for a distinguished state.
In addition to this value function that assigns a value to a state, there are also two additional functions that we will have to use. First, instead of assigning a value to a state, we could as well assign a value to a state and a given action. This action-value function is usually denoted by q and defined as
In other words, this is the expected discounted return if we start in state s and choose a as the first action before we start following the policy. Finally, the advantage function
essentially measure the additional benefit (or penalty) of choosing a as action in state s compared to the average under the policy.
PPO
To explain PPO, let us simplify things a bit and assume that we have picked a dedicated starting state s0 and are aiming at maximizing the outcome for this state only, i.e. we are trying to find a policy that maximizes the value of this state. Suppose that we start with a randomly chosen policy and are striving to identify a better policy
. As a starting point, we can relate the value of our state under the policy
to that under
and find, after some calculations that I will not reproduce here and that you can find in [1], that
Here, the expectation value is taken with respect to , more precisely it is taken with respect to the so-called discounted state distribution defined by
. We will not go through the proof, but intuitively, this makes sense – note that in order to maximize the right-hand side, we need to amplify the probabilities
for which the advantage taken with respect to
is positive, i.e. those actions which represent an improvement over the current policy
.
Let us now suppose that we model the policy using a neural network (spoiler: later, this will be our language model). We could then try to maximize the right hand side of this equation using gradient descent (or rather gradient ascent), and this relation would tell us that by doing so, we would obtain a better policy
. Unfortunately, there are two problems with this – the sampling and the unknown advantage function.
Let us first take a look at the advantage function. To calculate the loss function (or rather objective, as we want to maximize this, not minimize) we need the values of the advantage function. To do this, PPO uses the so-called actor-critic approach: in addition to the model describing the policy (called the actor model) which we want to optimize, we maintain a second model, called the critic, which is trained to learn the value function. We will later see that in each iteration of the PPO algorithm, we sample a few trajectories using the current policy, and during this phase, we can observe the resulting rewards and use them to derive an estimate for the advantage function. This estimate can then be used as ground truth for the critic model and we can apply gradient descent to train the critic model to approximate the advantage function as closely as possible. In practice, the actor model and the critic model have the same architecture and only differ in the top-level layer, or even share layers and weights.
The second problem is more difficult to solve. When looking at the equation above, we see that the policy – which we want to tune to maximize our objective – appears in two different places, namely inside the sum over the actions but also in the probability distribution that we use to define the expectation value. To estimate the expectation value, we would have to sample from this distribution, but after the first iteration of gradient ascent, we have changed
and all our samples, being taken with the previous value of the weights, would become stale. Even worse, sampling is a non-differentiable operation, so we cannot simply apply a backward pass to calculate the gradient that we need to apply to the weights.
In [1], an algorithm that has become known as TRPO (Trust region policy optimization) was proposed that deals with this problem by making an approximation. Specifically, instead of using the loss function (note the minus sign to be able to use gradient descent)
it uses the loss function
Note the subtle but important difference – we take the expectation value with respect to the distribution , not
as before. This approximation turns out to work well as long as
and
are not too different (in [1], this is spelled out using the so-called Kullback-Leibler divergence, we will see soon that PPO uses a slightly different approach to enforce this). Trivially, this is the same as
which we can write as an expectation value over the actions as well, namely
This is not yet exactly the loss function that PPO actually uses, but let us pause there for a moment and use this preliminary loss function to discuss the overall structure of the algorithm.

First, we initialize both models – the actor model that will learn our policy and the critic model that will learn the value function. Next, we perform the actual training iterations. In each iteration, we go through two phases as indicated in the diagram above – the sampling phase and the optimization phase.
In the sampling phase, we use our current policy to sample a few trajectories until we have collected data from a pre-defined number of state transitions, typically a few hundred or thousand. We do, however, not process these state transitions directly but first store them in a buffer.
One this is done, we enter the second phase, the optimization phase. In this phase, we randomly sample mini-batches from the previously filled buffer. Each item in one of the batches will give us one term of the form
i.e. one term in the loss function (behind the scenes, we use the critic model and a method known as GAE (see [2]) to calculate the advantage term). We add up these terms for a batch, obtain the loss function and apply a gradient descent step. This will give us new, updated weights of the actor model and therefore a new policy . Note, however, that the samples are still from the old policy
. We then repeat this for a given number of batches and epochs.
At this point, one iteration is complete. We now repeat the procedure for the next iteration, using as starting policy
from which we sample. In each iteration, we obtain a slightly improved policy , and we repeat this until we have reached convergence or a predefined number of iterations.
This already looks like a meaningful approach, but we have ignored one important point. Our loss function is an approximation, which gets worse within each iteration as starts to deviate significantly from
. To solve this, the PPO algorithm proposed in [3], uses the ratio
as a measure for the distance between and
. To control this distance and thus the error that we make by using the approximation for the loss function, PPO clips this ratio, i.e. restricts it to an interval of the form
where
is a hyperparameter that is often chosen to be 0.2.. This additional clipping yields the full loss function that PPO uses – in reality, the loss function is even a bit more complicated as the clipping logic depends on the sign of the advantage function and reads as follows.
PPO and language models
Having discussed PPO in a more general setting, let us now translate the terminology used in the previous few sections to the world of language models. First, we need to define what states and actions we will use to describe the training of a language model as a problem in reinforcement learning.
Intuitively, our state should correspond to the state of the conversation that a human has with the model. Therefore, a state is given by a prompt (which we would in practice sample from a set of prompts serving as test data) along with a partial completion, i.e. a sequence of token following the prompt. The actions are simply the token in the vocabulary, and the state transition is deterministic and modelled by simply appending the token. Here is an example.
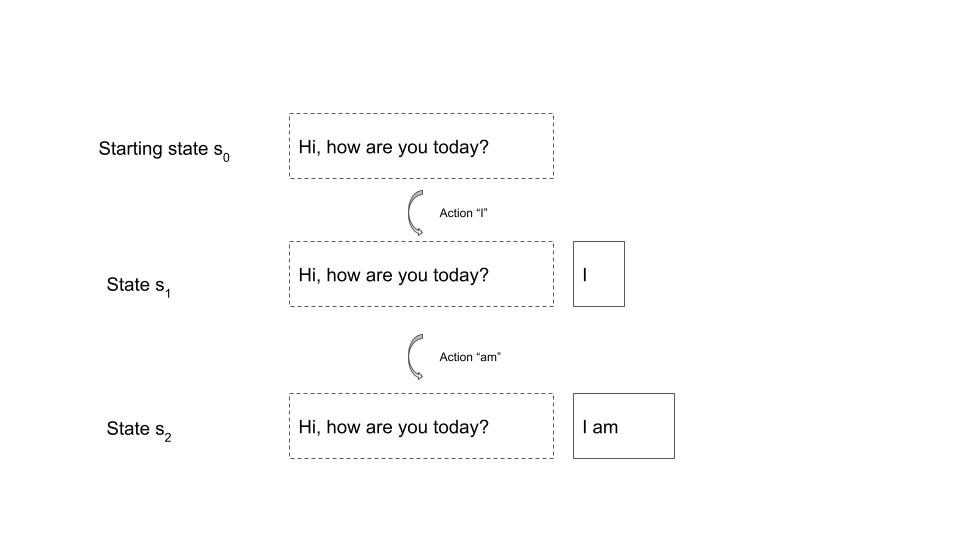
In this case, the initial state is the prompt “Hi, how are you today?”. When we sample a trajectory starting at this state, our first action could be the token “I”. We now append this token to the existing prompt and end up in the state given by the sequence of token “Hi, how are you today? I”. Next, we might sample the token “am” which takes us to the state “Hi, how are you today? I am” and so forth. A state is considered a terminal state if the last token is a dedicated end-of-sentence token used to indicate that the reply of the model is complete. Thus, an episode starts with a prompt and ends with a full reply from the model – in other words, an episode is a turn in the conversation with the model.
With this, it should also be clear what our policy is. A policy determines the probability for an action given a state, i.e. the probability of the next token given a sequence of token consisting of the initial prompt and the already sampled token – and this is of course what a language model does. So our policy is simply the language model we want to train.
To implement PPO, we also need a critic model. We could, for instance, again use a transformer that might or might not share some weights with the policy model, equipped with a single output head that models the value function.
Finally, we need a reward. Recall that the reward function is a (stochastic) function of states and actions. Thus, given an initial prompt x and a partial completion y, of which the last token is the selected action, we want to assign a reward r(x, y). This reward has two parts. The first part is only non-zero if y is complete, i.e. at the end of an episode, and – for reasons that will become clear in a minute – we will denote this part by
The second part is designed to address the issue that during reinforcement learning, the model might learn a policy that decreases the overall quality of the language model as it deviates too much from the initial model. To avoid this, this second term is a penalty based on the Kullback-Leibler divergence between the current policy and a reference policy (if you have never heard the term Kullback-Leibler divergence before, think of it as a distance between two probability distributions). This penalty term is
where is a hyperparameter which might be dynamically adapted during training.
Let us now talk about the actual reward RM(x, y). In an ideal world, this could be a reward determined by a human labeler that reflects whether the output is aligned with certain pre-defined criteria like accuracy, consistency or the absence of inappropriate language. Unfortunately, this labeling step would be an obvious bottleneck during training and would restrict us to a small number of test data items.
To solve this, OpenAI (building on previous work of other groups) trained a separate model, the so-called reward model, to predict the reward that a human labeler would assign to the reply. This reward model was first trained using labeled test data, i.e. episodes rated by a workforce of human labelers. Then, the output of this model (called RM(x,y) above) was used as part of the reward function for PPO. Thus, there are actually four models involved in the training process, as indicated in the diagram below.

First, there is the model that we train, also known as the actor model or the policy. This is the policy from which we sample, and we also use the probabilities that it outputs as inputs to the loss function. Next, there is, as always when using the PPO algorithm, the critic model which is only used during training and is trained to estimate the value of a state. The reward model acts as a substitute for a human labeler and assigns rewards to complete trajectories, i.e. episodes.
The reference model is a copy of the policy model which is frozen at the beginning of the training. This model is sometimes called the SFT model, where SFT stands for supervised fine tuning, as in the full training cycle used by OpenAI described nicely in this blog post, it is the result of a pretraining on a large data set plus fine tuning on a collection of prompts and gold standard replies created by a human workforce. Its outputs are used to calculate the Kullback-Leibler divergence between the current version of the policy and the reference model. All these ingredients are then put together in the loss function. From the loss function, gradients are derived and the critic as well as the actor are updated.
This completes our discussion of how PPO can be used to align language models with human feedback, an approach that has become known as reinforcement learning from human feedback (RLHF). Even though this is a long post, there are many details that we have not yet covered, but armed with this post and the references below, you should now be able to dig deeper if you want. If you are new to reinforcement learning, you might want to consult a few chapters from [6] which is considered a standard reference, even though I find the mathematics a bit sloppy at times, and then head over to [1] and [3] to learn more about PPO. If you have a background in reinforcement learning, you might find [3] as well as [4] and [5] a good read.
This post is also the last one in my series on large language models. I hope I could shed some light on the mathematics and practical methods behind this exciting field – if yes, you are of course invite to check out my blog once in a while where I will most likely continue to write about this and other topics from machine learning.
References
[1] J. Schulman et. al, Trust Region Policy Optimization, available as arXiv:1502.05477
[2] J. Schulman et al., High-Dimensional Continuous Control Using Generalized Advantage Estimation, available as arXiv:1506.02438
[3] J. Schulman et al., Proximal Policy Optimization Algorithms, available as arXiv:1707:06347
[4] D. Ziegler et al., Fine-Tuning Language Models from Human Preferences}, available as arXiv:1909.08593
[5] N. Stiennon et al., Learning to summarize from human feedback, available as arXiv:2009.01325
[6] R. S. Sutton, A.G. Barto, Reinforcement learning – an introduction, available online here